Project : step 3.4
Single-flagellated bacteria
(movement)
Disclaimer: the automatically generated English translation is provided only for convenience and it may contain wording flaws. The original French document must be taken as reference!
Goals : Simulate the specific movement of a first specific type of bacteria.
Our first concrete colony of bacteria will consist of bacteria known as single-flagellated bacteria.
These bacteria will largely retain the characteristics established in the Bacterium class but will be distinguished by a specific mode of movement (the move method).
Take the MonotrichousBacterium class that you were previously asked to draft and start by providing it with a constructor that takes only a value for its position as a parameter (a Vec2d). The other attributes will be initialized as follows:
- the energy level will be a random value uniformly distributed between getAppConfig()["monotrichous"]["energy"]["min"] and getAppConfig()["monotrichous"]["energy"]["max"].
The uniform function provided in Random.[hpp][cpp] allows you to generate such a random value.
- the direction will be a randomly generated unit Vec2d.
The provided function Vec2d::fromRandomAngle() allows you to generate such a vector.
- The radius will be a random value uniformly distributed between getAppConfig()["monotrichous"]["radius"]["min"] and getAppConfig()["monotrichous"]["radius"]["max"];
- and the color will be the value associated with getAppConfig()["monotrichous"]["color"].
For now, you don't need to worry about the mutable characteristics of the single-flagellated bacterium.
Specific Movement
Single-flagellated bacteria initially move in a random direction at a constant speed.
You are asked to code the movement method (move) for a single-flagellated bacterium according to the following algorithm:
- calculation of the new position and new speed (after a single time step has elapsed): see indications below
- decrease in energy level due to movement: energy -= length of movement * energy consumption factor (remember the consumeEnergy method)
- The distance traveled is the norm (method length) of the difference between the bacterium's new position and its old position (of the Vec2d). Note that the operator - is defined for Vec2d;
- the energy consumption factor is the parameter associated with ["energy"]["consumption factor"] (check your getters).
How do we calculate the new positions and velocities ?
As stated in the project description, we assume that the change in position and speed of a bacterium is governed by an equation of the form:

These methods require evaluating a force f for a given position, speed, and time.
For example, the Euler-Cromer integration scheme (see supplements, section 3.1) allows us to calculate:
new_speed = current_speed + dt * f(current_position, current_speed, t) new_position = current_position + dt * new_speed
There are other, more accurate integration schemes that operate on the same principle (notably the explicit Runge-Kutta algorithm).
The Euler-Cromer and Runge-Kutta methods are available in Utility/DiffEqSolver.[hpp] [cpp]: Calling the function stepDiffEq calculates the new position and speed of an object subjected to a force (referred to as eq in the parameters) based on its current position and speed, after a time interval of dt. The last parameter of the stepDiffEq function allows you to choose the integration method used (EC or RG4: for Euler-Cromer or Runge-Kutta of order 4). Both of these methods require evaluating the force (eq) for a given position and speed.
[Question Q3.18] The DiffEqFunction class (also found in the Utility/DiffEqSolver.hpp file) allows you to model a force f that is evaluated as a function of position and speed. Explain in your file REPONSES how you propose to use this class to provide a single-flagellated bacterium with a method that calculates the force f governing its movement? (hint: do we use composition or inheritance?)
Specify any attributes or methods, or any inheritance relationship, that you had to add to the class MonotrichousBacterium to perform the calculations required for the movement implementation, and justify your implementation choices.
Implement the method for calculating the force for a single-flagellated bacterium, noting that in this case ... the force is simply zero (zero two-dimensional vector, regardless of speed and position). This ensures that the bacterium moves in a uniform straight line.
To calculate the current speed, implement a method getSpeedVector that calculates the bacterium’s speed as its direction (unit vector) multiplied by a certain value (use the value 5; we’ll make this a bit more general later).
You now have all the information needed to implement the movement algorithm described above for single-flagellated bacteria. Integrate it into your program.
- Remember that a circular body has a move method.
- To avoid "jitter" effects in the movement, you can limit your circular body's movement to cases where the displacement calculated by stepDiffEq is not negligible: for example, when the squared norm (lengthSquared) of the displacement is greater than 0.001.
To avoid «running of the road»
With an accelerated simulation step (for example, a ["simulation"]["time"]["factor"] alue of 5), the bacterium may "exit" the culture dish before there is even time to perform the collision test! It would then be lost to the simulation. To avoid this problem, ensure that if the new position calculated by the move method places the bacterium outside the box, it also bounces back in the opposite direction.
Test 9 : Uniform linear motion
To test your MonotrichousBacterium class as it is currently coded, you will need to make it instantiable.Run the test using the target monotrichousTest. You should have uncommented it in the file CMakelists.txt and integrated this new material using Build > Run CMake.
Let a few nutrient sources generate, then create a single-flagellated bacterium (or several) using the 'M' key.
It must move straight ahead, consume nutrients in its path, and bounce off the edges of the culture dish as shown in the video below.
Improving the graphics
To improve the program's visual appearance, let's now add a sinusoidally moving flagellum to the rear of the single-flagellated bacterium.The colored circle used to represent it will therefore be supplemented by the drawing and animation of this flagellum.
Two aspects need to be addressed to implement this addition:
- drawing the animated flagellum;
- accounting for the bacterium’s change in direction as it moves, to ensure that the flagellum is always drawn at the rear.
Drawing the sinusoidal flagellum
The SFML library allows you to draw a set of points on a target graphics window using the following syntax :
auto set_of_points = sf::VertexArray(<type>); // we will discuss <type> a little later // adding points to the set: set_of_points.append(<point1>) ... set_of_points.append(<pointN>) target.draw(set_of_points);pointX can be specified in the form {{x, y}, color} where x and y are the coordinates of the point (of type float) and color is the SFML color used to color the point.
To draw a set of points corresponding to a sinusoidal flagellum, you must therefore fill in the set of points so that they form a sine wave.
The parameter <type> will be set to the type sf::PrimitiveType::TriangleStrip, which will allow the points in the set to be connected in pairs during rendering.
We will populate the set with absolute coordinates (centered at the origin of the coordinate system used by SFML). In the following section, we will apply a transformation to this set to place it in the correct position.
Here are some guidelines for populating the set of points so that they correspond to a sine wave with amplitude varying over time:
- add {0,0} as the first point to the set;
- For i from 1 to N (30 for example) : calculate the coordinates of the points to be added to the set as:
x = -i * radius / 10.f y = radius * sin(t) * sin(2 * i / 10.0)
where t is a double counter used to vary the amplitude of the sine wave over time. radius is the radius of the bacterium (the amplitude of the sine wave must be proportional to the graphical size of the bacterium).
- t can be initialized to a value uniformly distributed between 0 and PI and updated by adding 3 * dt.asSeconds() each time the bacterium moves.
- The coordinates x and y must be float; converting a double d to float can be done as follows: static_cast<float>(d).
[Question Q3.19] Where should the time variable t be declared and initialized ? Answer this question in your file REPONSES and implement the suggested changes.
At this point, we know how to fill in all the points of the sine wave in absolute coordinates. Before starting the drawing, we must place this set in the correct location, that is, behind the bacterium. Instructions are provided below.
Incorporating direction changes for the flagellum drawing
The Cartesian coordinate system used for SFML graphics rendering has its origin at the top-left corner of the graphics window.
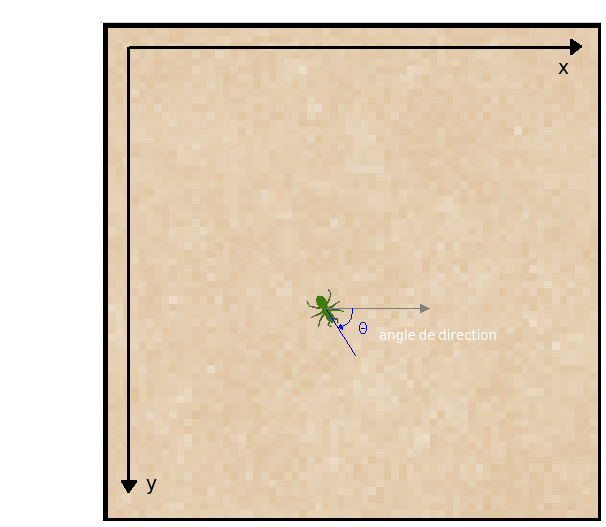
The direction of the bacterium’s movement is a vector pointing straight ahead (similar to the blue line pointing the insect forward in the figure below). We will call the angle formed by the bacterium’s direction vector relative to the x axis in this coordinate system (or in an orthogonal coordinate system centered on the bacterium, oriented like that of SFML) the direction angle.
|
SFML uses degrees, whereas C++ trigonometric functions use radians. You'll need to be careful about this.
|
To place the set of points in the correct location in the drawing, you can do the following:
auto transform = sf::Transform(); // declare a transformation matrix
// here, a set of operations such as translations or rotations applied to transform:
transform.translate(position);
rotate_and_translate(transform,
angle_direction / DEG_TO_RAD, // rotation
static_cast(-rayon + 2), // translation in x
0); // translation in y
// then:
target.draw(ensemble_de_points, transform); // Plot of all points
// after their transformation
// according to the transformation matrix
where position is the bacterium's position, radius is its radius as a circular path, and angle_direction is the direction angle (in radians) shown above.
This takes the direction angle into account to correctly position the flagellum at the rear of the bacterium.
Note that in the movement method, after a change in direction, the direction angle, expressed in radians, can be updated as follows :
auto const angleDiff = angleDelta(direction.angle(), rotation); // calculates the difference between the new
// direction angle and the old one
auto dalpha = PI * dt.asSeconds(); // calculates dα
dalpha = std::min(dalpha, std::abs(angleDiff)); // we cannot turn more than angleDiff
dalpha = std::copysign(dalpha, angleDiff); // rotate in the direction indicated by angleDiff
rotation += dalpha; // rotation angle updated
The functions/methodes angleDelta, angle and rotate_and_translate are given in Utility.[hpp][cpp] and Vec2d.[hpp][cpp].
Note that if dir is the direction of the bacterium, dir.angle() returns the rotation angle in radians associated with dir.
[Question Q3.20] How should you store the direction angle, and where in the code should you initialize and update it, assuming that all bacteria have a direction and a direction angle ? Answer this question in your file REPONSES and make the suggested additions to your code.
Test 10 : Uniform linear motion with flagellum animation
Run the test again as before; you should see your bacteria equipped with a splendid animated flagellum... which remains at the rear even when they bounce:
Back to the project description (Part 3)